Metabolism, Physiology, and Growth Characteristics of Bacilli
The Diversity of Bacilli
Previously, we studied the colonial, cellular, and chemical characteristics of a group of bacteria that had a common cellular morphology (they were cocci), but were quite diverse metabolically. To continue the investigation, we will use a similar approach to examine the cellular and metabolic characteristics of a second group of bacteria, this time all bacilli.
Bacilli show great diversity at the cellular level, beginning with the overall size and shape of the cells themselves. Some bacilli are short and plump little rods, while others are extremely slender and long. Curved and spiral shapes are common. A few species of bacilli produce endospores or other types of cellular inclusion bodies, such as metachromatic granules and parasporal crystals.
Bergey’s Manual initially divides the bacilli according to Gram stain reaction. Gram positive bacilli are further subdivided according to whether they form endospores, have filamentous growth or hyphae, and if they are acid-fast. Gram-negative bacilli are typically distinguished by size, shape, motility, and oxygen growth categories.
The group of bacteria included in this lab includes both Gram positive and Gram negative species. Using previous methods and additional tests, we will develop profiles of the bacilli as a way to distinguish among the various species.
Once again, although they are morphologically similar, the bacteria who have “volunteered” for this week’s lab differ in many ways and can be distinguished from each other by observing morphological and physiological characteristics. And like last week, we will begin by observing colonial and cellular morphology.
Morphological Evaluation of Bacterial Isolates
The bacteria we will examine in this lab include both Gram positive and Gram negative species of bacilli, each in a different genus. Observable differences start at the cellular level.
A volunteer from your lab bench should obtain cultures from your instructor, who will provide you with the species names. Write the name and BSL for each of the cultures below:
_________________________________________________________________
_________________________________________________________________
_________________________________________________________________
_________________________________________________________________
Colonial morphology
Distinguishing among bacterial species begins with examining colonial morphology, which you will do using the cultures listed below:
| Bacterium | Colonial Morphology |
| Bacillus_____________________
(write in Species name) |
|
| Corynebacterium______________
(write in Species name) |
|
| Pseudomonas aeruginosa |
Gram stain and cellular morphology
Among the bacteria in this group, there are significant observable differences in cellular morphology that can be observed by staining the cells, beginning with the Gram stain. For each culture listed, Gram stain a smear you prepare, and record the Gram stain reaction, cellular morphology, and arrangement. Also if you observe any other cellular features, specifically endospores or inclusion bodies in the Gram stained smear, include those observations in your description as well.
| Bacterium | Gram stain results |
| Bacillus _____________________ (write in Species name) | |
| Corynebacterium _____________
(Write in Species name) |
|
| Pseudomonas aeruginosa |
A major distinction among Gram positive bacilli is the production of endospores, which can be observed by endospore staining. For the two Gram positive species of bacteria, prepare a second smear and stain it using the endospore stain method. If endospores are observed, report the shape and location.
| Bacterium | Endospores present? (if yes, describe) |
| Bacillus______________________ (write in Species name) | |
| Corynebacterium______________ (write in Species name) |
Physiology and growth characteristics
As previously done with the cultures of cocci, evaluate energy metabolism and physiological oxygen requirements for the bacteria in the table below by performing the catalase, oxidase, Triple Sugar Iron (TSI), and nitrate reduction tests.
|
Bacterium |
Catalase |
Oxidase |
Nitrate Reduction |
TSI |
| Bacillus __________________
(write in Species name) |
||||
| Pseudomonas aeruginosa | ||||
| Salmonella________________
(write in Species name) |
From the outcome of the tests above, determine if there is evidence of aerobic or anaerobic respiration or fermentation, and indicate why you reached that conclusion.
| Bacillus ________________ | State what evidence from test results indicates the bacteria use this pathway. If there is no evidence, write “none.” |
| Aerobic respiration | |
| Anaerobic respiration | |
| Fermentation |
Oxygen Growth Requirement Category____________________________________________
| Pseudomonas aeruginosa | State what evidence from test results indicates the bacteria use this pathway. If there is no evidence, write “none.” |
| Aerobic respiration | |
| Anaerobic respiration | |
| Fermentation |
Oxygen Growth Requirement Category____________________________________________
| Salmonella _____________ | State what evidence from test results indicates the bacteria use this pathway. If there is no evidence, write “none.” |
| Aerobic respiration | |
| Anaerobic respiration | |
| Fermentation |
Oxygen Growth Requirement Category____________________________________________
Differentiating Among Bacterial Species
In a clinical laboratory, differentiating the nonpathogenic microbiota found in the GI tract from pathogenic species has historically relied on phenotypic tests. The small and large intestines are home to the largest number of bacterial species, which contribute greatly to human health and disease. Generally, Gram positive cocci and bacilli are considered “probiotic” because of the health promoting benefits associated with these bacteria, while Gram negative species are commensals that can induce inflammation if their proliferation is not kept in check.
Many of the Gram negative bacteria residing in the GI tract are members of the Family Enterobacteriaceae. Bacteria in this family are facultative anaerobes that are usually oxidase-negative and ferment glucose, which distinguishes them from aerobic species of Gram negative rods. To differentiate among them, a set of tests with the acronym IMViC, which stands for: Indole, Methyl red, Voges-Proskauer, and Citrate (with the lower-case “I” thrown in to make it easier to say). The TSI and urease tests and others may also be performed at the same time to improve the identification process.
Examples of IMViC results for some of the Enterobacteriaceae, which illustrate how these tests can be used to differentiate among the bacteria in this group, are given in Table 1. As evidenced by the results shown for the two species of Proteus, metabolic differences may even exist among individual species within the same bacterial genus.
| Table 1 | ||||
| Species | Indole | Methyl Red | Voges-Proskauer | Citrate |
| Escherichia coli | Positive | Positive | Negative | Negative |
| Shigella spp. | Negative | Positive | Negative | Negative |
| Klebsiella spp. | Negative | Negative | Positive | Positive |
| Proteus vulgaris | Positive | Positive | Negative | Negative |
| Proteus mirabilis | Negative | Positive | Negative | Positive |
Sulfide–Indole–Motility (SIM)
This is another medium in which a number of different reactions may occur. In fact, this medium is used to determine if (1) the bacterium reduces sulfate and produces H2S, which is evidence of anaerobic respiration; (2) the culture oxidizes the amino acid tryptophan and produces indole, and (3) the bacterium is motile. The medium contains sulfur (as the oxidizing agent) and ferrous ions (for H2S detection), tryptophan (for indole detection), and is prepared in tubes referred to as stabs or deeps (because they are not slanted). The medium also has a lower concentration of agar and is semi-solid, to detect bacterial motility. Bacteria are inoculated in this medium with an inoculating needle (as opposed to a loop) and stabbing the bacteria deep into the soft agar.
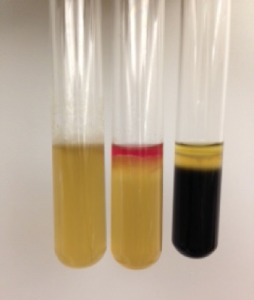
After incubation, a positive test for sulfide turns the media black, just as for the TSI test. Tryptophan oxidation by the bacteria yields indole, which can be detected by adding a reactive chemical, which turns deep pink. Motility, which may be the hardest to determine, is noted by looking closely at the stab line. Motile bacteria swim through the semi-solid agar and the medium appears cloudy, obscuring the stab line. If the bacteria are not motile, the stab line is clearly visible. The outcomes for all three tests (sulfide, indole, and motility) should be noted. Figure 1 shows three SIM tubes, each showing different outcomes for the three tests. From left to right, the results are:
(1) Sulfide negative, Indole negative, Motility positive
(2) Sulfide negative, Indole positive, Motility positive
(3) Sulfide positive, Indole negative, and Motility positive
How would you interpret the results for the bacteria inoculated in the SIM medium shown below?

Sulfide: _____________________________________
Indole: ______________________________________
Motility: _____________________________________
Methyl Red and Voges-Proskauer (MR-VP)
Fermentation of glucose by bacteria may produce one or more acids or alcohols. Probiotic lactic acid (homolactic) bacteria produce lactic acid as the major end product. Other fermentation pathways yield a combination of acids and/or alcohols. The organic end products of these pathways can be detected using chemical methods. The medium used for these tests, MR-VP broth is the same for both tests. The broth contains glucose as a fermentable substrate, and the end products are measured after incubation.
Methyl Red (MR)
Bacteria utilizing a mixed acid fermentation produce stable organic acid end products. After the bacteria grow, the pH indicator methyl red (which is red below pH 4) is added to the culture. If the broth is red after methyl red is added, the result is considered a positive test. For bacteria that produce alcohols and other less acidic metabolites, the indicator (and therefore the medium) turns orange to yellow, which is a negative result (see Figure 2a)
Voges-Proskauer (VP)
Some bacteria use a butanediol fermentation pathway when glucose is the fermented substrate. The end products of this type of fermentation include a variety of both acidic and non-acidic end products, including ethanol, butanediol, and acetoin. If acetoin is present, chemical reagents added to the broth will react with it and form a reddish-brown colored compound, which is considered a positive test result (see Figure 2b).


Citrate utilization
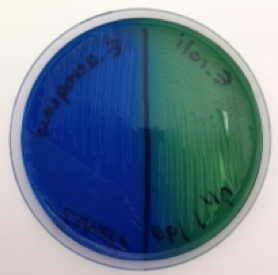
Some bacteria have an enzyme (citrate permease) that facilitates the transport of citric acid into the cell, and another enzyme, citrase, for citrate catabolism. These bacteria have the ability to grow on a medium (Simmons’ citrate agar) that contains nothing more than citric acid, the first intermediate in the Krebs cycle of aerobic respiration, as a food source. Metabolism of citric acid releases carbon dioxide, which reacts with sodium and water in the medium to form a compound with a basic (alkaline) pH. In the presence of a base, the pH indicator in the medium (bromothymol blue) changes color from green to blue, which is a positive result for this test. The Simmons’ citrate agar plate shown in Figure 3 illustrates both a positive (on the left) and negative (on the right) result for this test.
| Table 2. Summary of Test Methods | |
| Test | Method |
| SIM | With an inoculating loop, stab culture to the bottom of a SIM deep tube; incubate. AFTER INCUBATION, add 5 drops of indole (Kovac’s) reagent and observe for deep red color. |
| Methyl Red (MR) | Transfer 2 ml of MR-VP broth to a sterile culture tube and inoculate with bacteria; incubate. AFTER INCUBATION, add 5 drops of methyl red reagent and observe for red color. |
| Voges–Proskauer (VP) | Transfer 2 ml of MR-VP broth to a sterile culture tube and inoculate with bacteria; incubate. AFTER INCUBATION, add 5-10 drops of Reagent A followed by 5-10 drops of Reagent B. Allow the tube to sit for at least 20 minutes and observe for red color. |
| Citrate | Inoculate the surface of a Simmons’ citrate agar plate and incubate. Observe for blue color change. |
After the tests have been demonstrated, perform them on the four Enterobacteriaceae listed in the table below.
| Species | Indole | Methyl Red | Voges-Proskauer | Citrate |
| Escherichia coli | ||||
| Salmonella spp. | ||||
| Citrobacter freundeii | ||||
| Enterobacter aerogenes |
Construction of a dichotomous key to differentiate among this group of bacteria
In the space below, construct a dichotomous key to show how the tests you performed may be used to identify ALL of the different bacilli we used in this lab.

